Abstract
To better understand how amino acid sequence encodes protein structure, we engineered mutational pathways that connect three common folds (3α, β−grasp, and α/β−plait). The structures of proteins at high sequence-identity intersections in the pathways (nodes) were determined using NMR spectroscopy and analyzed for stability and function. To generate nodes, the amino acid sequence encoding a smaller fold is embedded in the structure of an ~50% larger fold and a new sequence compatible with two sets of native interactions is designed. This generates protein pairs with a 3α or β−grasp fold in the smaller form but an α/β−plait fold in the larger form. Further, embedding smaller antagonistic folds creates critical states in the larger folds such that single amino acid substitutions can switch both their fold and function. The results help explain the underlying ambiguity in the protein folding code and show that new protein structures can evolve via abrupt fold switching.
Introduction
There have been remarkable advances recently in the ability to predict the tertiary structure of a protein from its primary amino acid sequence1,2as well as to design amino acid sequences that encode stable, unique protein structures3。It is also well-established, however, that some proteins have a propensity for two completely different, but well-ordered, conformations4,5,6,7,8,9,10,11,12。Better insight into the ambiguity of the protein folding code would lead to a better understanding of how proteins evolve, how mutation is related to disease, and how function can be annotated to sequences of unknown structure13,14,15,16,17,18,19,20,21,22,23,24,25,26,27。If the protein folding code were truly understood, it would be possible both to predict and design proteins that undergo profound switches in conformation. There has been significant progress in understanding natural proteins that switch folds11and predicting natural fold-switching proteins from amino acid sequence data25。Designing proteins at the interface between different folds has been possible7,28,29,30but still presents a formidable challenge. It has been particularly challenging to design monomeric proteins that switch fold without a change in quaternary structure, and a better understanding is needed about how a very limited subset of intra-protein interactions can tip the balance from one fold and function to another29,31,32。
Our goal here was to engineer monomeric proteins that are in a critical state between two distinct folds. To do this we chose three well-studied protein folds and designed a series of sequences such that each sequence is compatible with two sets of native interactions. Two of these folds are fromStreptococcalProtein G which contains two types of domains that bind to serum proteins in blood: the GAdomain binds to human serum albumin (HSA)33,34和克Bdomain binds to the constant (Fc) region of IgG35,36。The third protein is S6, a component of the 30S ribosomal subunit ofThermus thermophilus37,38,39,40,41。For simplicity, the S6 fold is referred to as an S-fold, the GAfold as an A-fold, and the GBfold as a B-fold. These proteins share no significant sequence homology and are representative of three of the ten most common folds: the S-fold is a thioredoxin-like α/β plait; the A-fold is a homeodomain-like 3α-helix bundle; and the B-fold is a ubiquitin-like β grasp42。
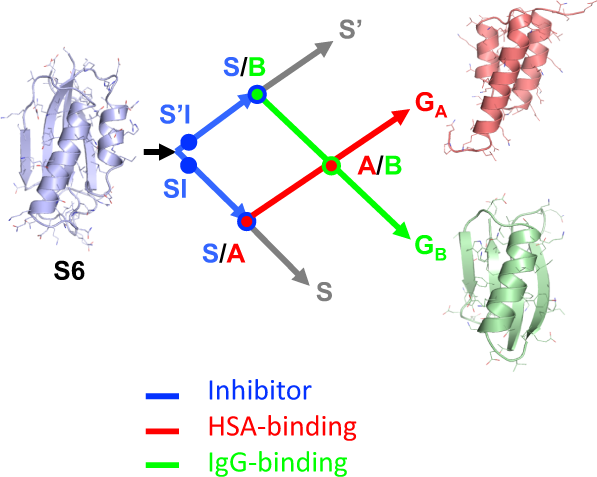
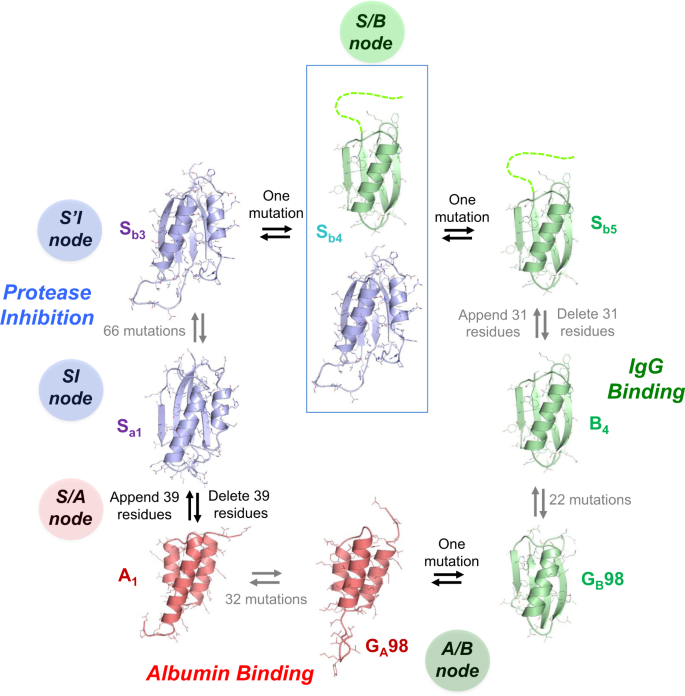
Figure1depicts a network of high-identity sequence intersections (nodes) that connect the three folds. The arrows in Fig.1show a network originating with the natural S6 sequence. Circles represent nodes in the network at which structural and/or functional switches occur. The SI and S’I nodes are branch points and lead down diverging sequence pathways, one leading to a node with the A-fold (S/A) and one to a node with the B-fold (S/B). Intersecting mutational pathways lead from S/A to the native GAprotein and S/B to the native GBprotein. At this intersection (A/B), an A-fold switches to a B-fold.
S6是原点equence in the engineering process. SI and S’I are separate nodes and are loop-size variants of the S-fold, both having protease inhibitor functions. The SI branch of the mutational path leads to a node with the A-fold and HSA binding function. The S’I branch of the path leads to a node with the B-fold and IgG binding function. The S/A node (blue and red circles) includes proteins Sa1, Sa2, A1, and A2。The S/B node (blue and green circle) includes proteins Sb3, Sb4, Sb5, B3, and B4。The A and B paths themselves intersect at an A/B node (green and red circles) at which A- and B-folds are nearly iso-energetic and bifunctional. The S and S’ branches continue and connect with many other natural sequences in the α/β plait super-fold family.
Proteins around the A/B node have been extensively characterized in our earlier work29,31,32。Here we determine that both GAand GBcan switch into a third fold (α/β−plait) and show that these three folds and four functions (HSA-binding, IgG-binding, protease inhibition, and RNA-binding) can be connected in a network that avoids unfolded and functionless states. We describe how these nodes were engineered, determine key structures using NMR spectroscopy, and analyze stability and binding function. The ability to design and characterize nodes connecting three common small folds suggests that fold switching may be an intrinsic feature of the protein folding code and is important in the evolution of protein structure and function.
Results
Designing a functional switch from ribosomal protein to protease inhibitor
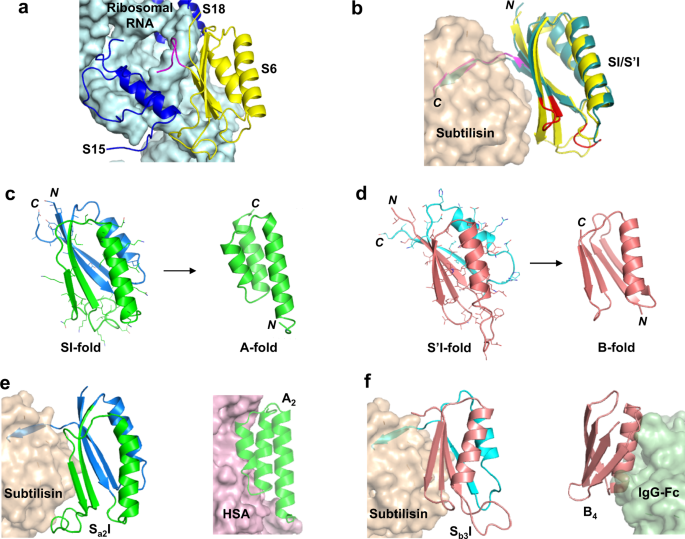
The S6 ribosomal protein is structurally homologous to subtilisin protease inhibitors known as prodomains (Fig.2a, b)43,44。Prodomain-type inhibitors have two binding surfaces with the protease. One surface comprises the last nine C-terminal amino acids of the inhibitor which bind in the substrate binding cleft of the protease (Fig.2b).A second, more dynamic surface is formed between two subtilisin helices and the large surface of the β−sheet in the α/β-plait topology of the inhibitor (Fig.2b)45,46,47。As a result, the S6 protein could be converted into a subtilisin inhibitor protein of the same overall fold (denoted SI) by replacing its nine C-terminal amino acids with residues optimized to bind in the substrate binding cleft of subtilisin. This replacement results in new contacts between the SI β−sheet and the subtilisin surface helices (Fig.2b).
aStructure of the S6 protein (yellow), RNA (light blue), and S15 and S18 proteins (blue) in the 30S ribosome (PDB 1FKA [https://doi.org/10.2210/pdb1FKA/pdb], ref.43).C-terminal amino acids of S6 are in magenta.bSubtilisin (wheat) is shown in complex with a model of the SI-inhibitor (yellow). The C-terminal nine amino acids of SI are shown in magenta. These positions were mutated in native S6 to generate affinity for subtilisin. The S’I-inhibitor (teal) is also shown with the altered loops in red. The subtilisin used in the modeling was the engineered RAS-specific protease.cThe Sa1protein (blue and green) was generated from SI by mutating the 45 positions (mutant side chains shown with sticks). Deletion of C-terminal amino acids (blue) switches Sa1into the A-fold (green).dThe Sb3protein (rose and cyan) was generated from S’I by mutating the 67 positions (mutant side chains shown with sticks). Deletion of C-terminal amino acids (cyan) or point mutation will switch Sb3into a B-fold (rose).eModel of Sa2I (green and blue) bound to subtilisin (wheat). Model of A2(基于1圣ructure) bound to HSA (violet). The HSA complex usedPDB 2VDB(ref.50) as the template.fModel of Sb3(rose and cyan) bound to subtilisin (wheat). Model of B4(rose) bound to Fc (mint). The Fc complex usedPDB 1FCC(ref.51) as the template. The subtilisin used in the modeling and inhibition measurements was the engineered RAS-specific proteasePDB 6UAO(ref.49).
The SI-protein is 99 amino acids in length and has a 10 residue loop between β2 and β3. However, there are many natural variations in the length of loops in the conserved α/β-plait topology48。Therefore, we also engineered a 91 amino acid version of the S-fold (denoted S’I), which resembles the topology of natural prodomain inhibitors (Supplementary Fig.1).Specifically, the S’I inhibitor has a longer loop connecting β1 to α1 and a shorter turn connecting β2 to β3 (Fig.2b).
The SI and S’I proteins were expressed and purified by binding to a protease column49。The CD spectra were compared to the native S6 protein (Supplementary Fig.1).Inhibition constants (KI) were measured using an engineered RAS-specific subtilisin protease and the peptide substrate QEEYSAM-AMC49。SI and S’I inhibit the RAS-specific protease withKIvalues of 200 and 60 nM, respectively (Supplementary Table1).竞争性抑制实验的细节are described in the “Methods” section. The results demonstrate that a ribosomal protein can be converted into a protease inhibitor with minor modification (and without a fold switch). In addition, however, the SI and S’I proteins also facilitated engineering subsequent switches to new folds and functions by linking each of the S-, A-, and B- folds to easily measured binding functions: protease inhibition (S or S’-fold); HSA-binding (A-fold, Fig.2e)50; and IgG binding (B-fold, Fig.2f)51。
Designing fold switches
In previous work, we created sequences that populate both A- and B-folds by threading the A-sequence through the B-fold, finding a promising alignment, and then using phage-display selection to reconcile one sequence to both folds29,52,53。Here the approach is conceptually similar, except that we use Rosetta54as a computational design tool to test compatible mutations rather than phage display. The design process is as follows:
- i.
Thread the A- or B- sequence through both SI and S’I-fold types.
- ii.
Identify alignments that minimize the number of catastrophic interactions.
- iii.
Design mutations to resolve unfavorable interactions in clusters of 4–6 amino acids using Pymol55and energy minimize using Rosetta-Relax54。
- iv.
Optimize protein stability in the S-fold by computationally mutating amino acids at non-overlapping positions. Repeat energy minimization and evaluation with Rosetta-Relax.
- v.
To reduce uncertainties involved in computational design, conserve original amino acids whenever possible.
There is no reason to assume that this method is optimal. We are just applying a practicable scheme for engineering sequences compatible with two sets of native interactions and then evaluating structure, stability, and function. Initial designs were refined based on structural analysis with NMR, thermodynamic analysis of unfolding, and functional analysis using binding assays, as described below. All designed proteins were expressed inE。coliand purified to homogeneity as described in the “Methods” section.
Designing a switch from α/β-plait protease inhibitor to 3α HSA-binding protein
Alignment of the 56 amino acid HSA-binding, A-fold with the 99 amino acid SI-fold and subsequent mutation to resolve catastrophic interactions produced low-energy switch candidates denoted Sa1and A1。The exact sequence of A1is embedded in Sa1at positions 11–66 such that the α1 helices are structurally aligned (Fig.3a, Supplementary Fig.2A).Their final computational models were generated by Rosetta using the Relax application. The Relax protocol searches the local conformational space around an experimentally determined structure and is used only to evaluate whether the designed mutations have favorable native interactions within that limited conformational space. The designed models of Sa1and A1are very similar in energy compared to the respective relaxed native structures (Supplementary Fig.3and Source data files).
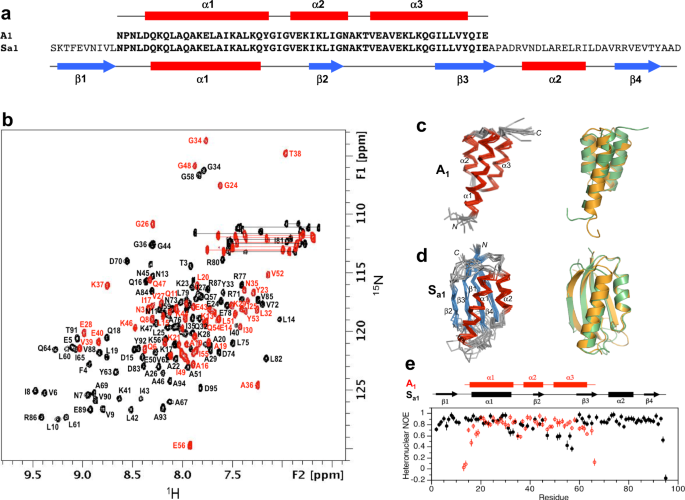
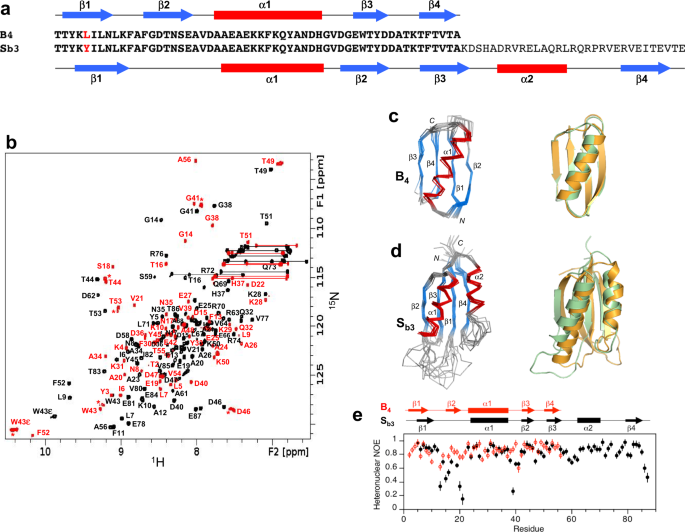
aSequence alignment of A1and Sa1, which are 100% identical over the 56 amino acid A-region.bOverlaid two-dimensional1H-15N HSQC spectra of Sa1(black) and A1(red) with backbone amide assignments. Spectra were recorded at 25 and 5 °C, respectively.cEnsemble of 10 lowest energy CS-Rosetta structures for A1(left panel). Superposition of the A1圣ructure (green) with the parent GAfold (orange) (right panel).dEnsemble of 10 lowest energy CS-Rosetta structures for Sa1(left panel). Superposition of Sa1(green) with the parent S6 fold (orange) (right panel).eBackbone dynamics in designed proteins. Plot of {1H}-15N steady state heteronuclear NOE values at 600 MHz versus residue for A1(red) and for Sa1(black). Each set of heteronuclear NOEs was obtained from a single experiment. Errors were estimated based on the measured background noise level.
Structural analysis of A1and Sa1
Overall, the 3α-helical bundle topology of A1is very similar to the GAparent structure from which it was derived56。The sequence-specific chemical shift assignments for A1(Fig.3b) were utilized to calculate a 3D structure with CS-Rosetta (Fig.3c, Table1).Our previous studies indicated close correspondence of CS-Rosetta and de novo structures for A- and B-folds with high sequence identity57。The N-terminal residues 1–4 and the C-terminal residues 53–56 are disordered in the structure, consistent with {1H}-15N steady-state heteronuclear NOE data (Fig.3e).Likewise, Sa1has the same overall βαββαβ-topology as the parent S6 structure (Fig.3d, Table2).The backbone chemical shifts (Fig.3b) were used in combination with main chain inter-proton NOEs (Supplementary Fig.4) to determine a three-dimensional structure utilizing CS-Rosetta (PDB 7MN1).The conformational ensemble shows well-defined elements of secondary structure at residues 2–10 (β1), 16–32 (α1), 40–44 (β2), 59–67 (β3), 73–81 (α2), and 86–92 (β4). The principal difference from the native structure is that the β2-strand is seven amino acids shorter in Sa1than in S6. Heteronuclear NOE data show overall consistency with the structure, indicating that the long loop between the β2- and β3-strands from residues 45–58 is more flexible than other internal regions of the polypeptide chain (Fig.3e).
Comparison of A1and Sa1圣ructures
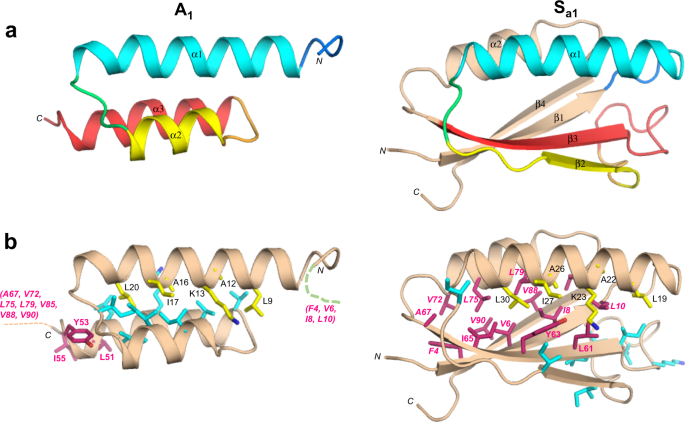
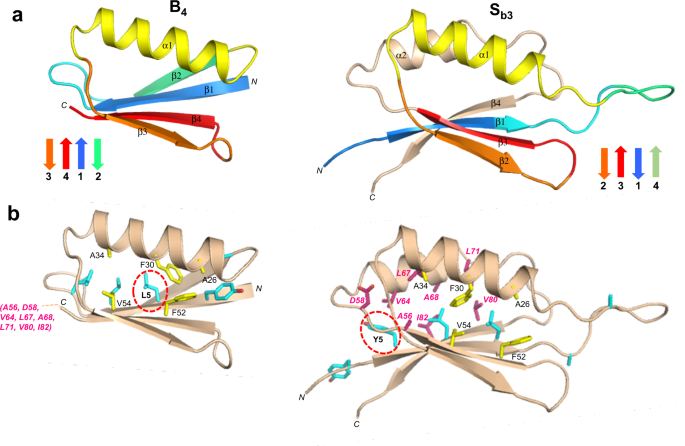
Although the 56 amino acid sequence of A1is 100% identical to residues 11–66 of Sa1, a significant fraction of the backbone undergoes changes between the two structures. Most notably, while the α1 helices in both A1and Sa1are similar in length, the regions corresponding to the α2 and α3 helices of A1form the β2 and β3 strands of Sa1(Fig.4a).Core amino acids in the α1-helix of A1与残留,也导致the core of Sa1。However, the α1-helix in Sa1contacts an almost entirely different set of residues (Fig.4b).For example, amino acids L51, Y53, and I55 in the C-terminal tail of A1do not have extensive contact with α1 but the corresponding residues in Sa1(L61, Y63, and I65) form close core interactions with α1 as part of the β3-strand. Most of the other core residues contacting the α1-helix of Sa1are outside the 56 amino acid region coding for the A1褶皱。这些包括F4, V6, I8 L10β1-strand; A67 from the β3-strand; V72, L75, and L79 from the α2-helix; and V85 from the loop between the α2-helix and the β4-strand. Two additional residues, V88 and V90 (β4) also contribute significantly to the core but do not contact α1. Thus, except for the original topological alignment of the α1-helices, the cores of the 3α and α/β-plait folds are largely non-overlapping. In total, approximately half of the residues participating in the Sa1core are not present in the A1sequence.
aMain chain comparisons. (Left panel) CS-Rosetta structure of A1with color coding for secondary structured elements. (Right panel) Corresponding color-coded regions mapped onto the CS-Rosetta structure of Sa1, illustrating changes in backbone conformation. Regions outside the 56 amino acid sequence of A1are shown in wheat.bSide chain comparisons. (Left panel) Residues contributing to the core of A1from the α1-helix (yellow), and from other regions (cyan). The non-α1 core residues from Sa1(pink) do not overlap with the A1core (see text for further details). (Right panel) Residues contributing to the core of Sa1from the α1-helix (yellow), and most of the other participating core residues (pink). The non-α1 core residues from A1are also shown (cyan), highlighting the low degree of overlap.
Energetics of unfolding for A1/Sa1
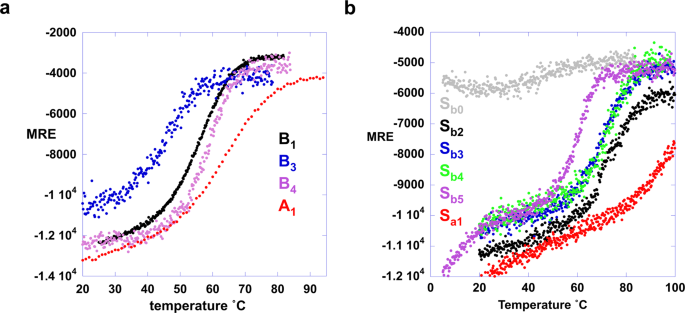
Far-UV CD spectra were measured for Sa1and A1and their thermal unfolding profiles were determined by measuring ellipticity at 222 nm versus temperature (Fig.5and Supplementary Fig.5).Sa1has a TMof ~100 °C and an estimated ∆Gfoldingof −5.3 kcal/mol at 25 °C (Fig.5b, Supplementary Table1)58。The ∆Gfoldingof the parent S6 is −8.5 kcal/mol40。The Rosetta energy of the Sa1design is similar to that of the native sequence (Supplementary Fig.3).A1has aTMof 65 °C and a ∆Gfolding = −4.0 kcal/mol at 25 °C58(Fig.5a, Supplementary Table1).The ∆Gfoldingof the parentGAis −5.6 kcal/mol59,60。The Rosetta energy of the A1design is slightly more favorable than for the native sequence (Supplementary Fig.3).
HSA binding
Initial engineering of the fold switch was carried out without consideration of preserving function. As a result, A1does not have detectible HSA binding affinity because two amino acids in the binding interface were mutated. Significant HSA-binding is recovered, however, when the surface mutations, E28Y and K29Y, are made in A1(denoted A2).These mutations do not appear to affect the structure of A1(Supplementary Fig.5) but result in HSA binding ofKD ≤ 1 µM (Supplementary Table1).This was determined by measuring binding to immobilized HSA as described in the “Methods” section.
Protease inhibition
Sa1does not bind protease because C-terminal amino acids were not preserved in its design. It can be converted into a protease inhibitor, however, by replacing its three C-terminal amino acids (AAD) with DKLYRAL (denoted Sa1I). A version of Sa1I was also made that contains the exact 56 amino acid A2sequence by making E38Y, K39Y mutations (denoted Sa2I). Sa1, Sa1我和年代a2I are similar in structure by CD analysis (Supplementary Fig.5).The inhibition constant of Sa2I with the engineered subtilisin was determined to be 50 nM as described in the “Methods” section (Supplementary Table1).Thus, a stable A-fold with HSA-binding function can be embedded within a 99 amino acid S-fold with protease inhibitor function (Fig.2c, e).It should be noted that all HSA contact amino acids are preserved in both the A2and Sa2I sequences, but the three-dimensional topology necessary to form the HSA contact surface occurs only in the A-fold50。Nevertheless, Sa2I was observed to bind weakly to HSA (KD ~ 100 µM, Supplementary Table1).This weak affinity suggests that some Sa2I molecules may populate the 3α fold even though the α/β-plait fold strongly predominates.
Designing a switch from α/β-plait protease inhibitor to β−grasp IgG-binding protein
In designing an S- to B-fold switch, we used two topological alignments. The first was between SI- and B-folds, where the β1 strands of each fold were aligned (Supplementary Figs.2B and6A).The second alignment was between S’I- and B-folds, where the long loop between β2 and β3 in SI was shortened in S’I to be more consistent with natural protease inhibitors. In this scheme, the α1β3β4 topology of the B-fold was aligned with the α1β2β3 topology of the S’I-fold (Fig.6a, Supplementary Fig.2C).
aSequence alignment of B4and Sb3, differing by one residue (L5Y) over the 56 amino acid B-region.bOverlaid two-dimensional1H-15N HSQC spectra of Sb3(black) and B4(red) with backbone amide assignments. Spectra were recorded at 25 °C. The A56 peak is an aliased signal. Peaks labeled with an asterisk decrease in relative intensity as the B4concentration is lowered, indicating the presence of a weakly associated putative dimer in addition to monomer.cEnsemble of 10 lowest energy CS-Rosetta structures for B4(left panel). Superposition of the B4圣ructure (green) with the parent GBfold (orange) (right panel).dEnsemble of 10 lowest energy CS-Rosetta structures for Sb3(left panel). Superposition of Sb3(green) with the parent S6 fold (orange) (right panel).ePlot of {1H}-15N steady-state heteronuclear NOE values at 600 MHz versus residue for B4(red) and Sb3(black). Each set of heteronuclear NOEs was obtained from a single experiment. Errors were estimated based on the measured background noise level.
Design and characterization of B1, Sb1, B2, and Sb2
In the first approach, alignment of the β1-strands of the B-fold and the S-fold and subsequent mutation to resolve catastrophic interactions produced low-energy switch candidates denoted B1and Sb1。The exact sequence of B1is embedded in Sb1at positions 4–59 (Supplementary Fig.6A).The computational models of B1and Sb1show relatively small increases in energy compared to the corresponding relaxed native structures (Supplementary Fig.3).The NMR structure of B1displayed a ββαββ topology identical to that of the parent B-fold, with a backbone RMSD of ~0.6 Å (Supplementary Fig.6B, C).The topology of Sb1is not the same as the parent S6 structure, however, and instead has a fold similar to that of B1(Supplementary Figs.6B, D, and7,PDB 7MQ4).Introducing 13 mutations into Sb1generated a protein denoted Sb2(Supplementary Fig.8).Sb2contains four β-strands and two α-helices and has the general features of the parent S-fold (Supplementary Fig.9,PDB 7MN2).The 56 amino acid version of Sb2(denoted B2) has a significantly higher Rosetta energy than B1, however, and is presumably unfolded (Supplementary Fig.3).Thus, neither the B1/Sb1nor B2/Sb2protein pairs resulted in high identity sequences with different folds. Nonetheless, B1is 80% identical to the corresponding embedded region in the S-folded protein Sb2(Supplementary Fig.9A).The structures of B1, Sb1, and Sb2are described further in the Supplement and Tables1and2。
Design of Sb3and B3
To improve the design of the S-to-B switch we aligned the B-fold with the S’ inhibitor fold and chose an alignment that creates a topological match between α1β3β4 in B and α1β2β3 in S’ (Supplementary Fig.2C).Mutation to resolve deleterious interactions in this alignment produced low-energy switch candidates denoted B3and Sb3(Supplementary Fig.10).The exact sequence of B3is embedded in Sb3at positions 1–56. The energy of the computational model for Sb3is slightly more favorable than the relaxed native structure. The designed model of B3shows relatively small increases in energy compared to the relaxed native structure (Supplementary Fig.3).
Structural analysis of Sb3and B3
NMR-based structure determination indicated that Sb3has a βαββαβ secondary structure and an S-fold topology (Fig.6a, b, d,PDB 7MP7).Ordered regions correspond with residues 4–10 (β1), 24–37 (α1), 42–46 (β2), 51–56 (β3), 62–70 (α2), and 79–85 (β4). Comparison of Sb3with the parent S-fold indicates that the β1/α2/β4 portion of the fold is similar in both. In contrast, the β1–α1 loop is longer in Sb3(13 residues) than in the parent S-fold (5 residues), while α1, β2, the β2–β3 loop, and β3 are all shorter than in the parent (Fig.6d).Consistent with the Sb3圣ructure, the 13 amino acid β1-α1 loop is highly flexible (Fig.6e).We also expressed and purified a truncated protein corresponding to the embedded B-fold, the 56 amino acid version of Sb3(denoted B3).The 2D1H–15N HSQC spectrum of B3at 5 °C and low concentrations (<20 μM) was consistent with a predominant, monomeric B-fold (Supplementary Fig.11) but showed significant exchange broadening at 25 °C, indicative of low stability (see below). Presumably, the low stability is due to the less favorable packing of Y5 in the core of the B-fold compared with a smaller aliphatic leucine. However, additional, putatively oligomeric, species were also present for which relative peak intensities increased with increasing protein concentration. Due to its relatively low stability and sample heterogeneity, B3was not analyzed further structurally.
Design and analysis of point mutations that switch the fold of Sb3
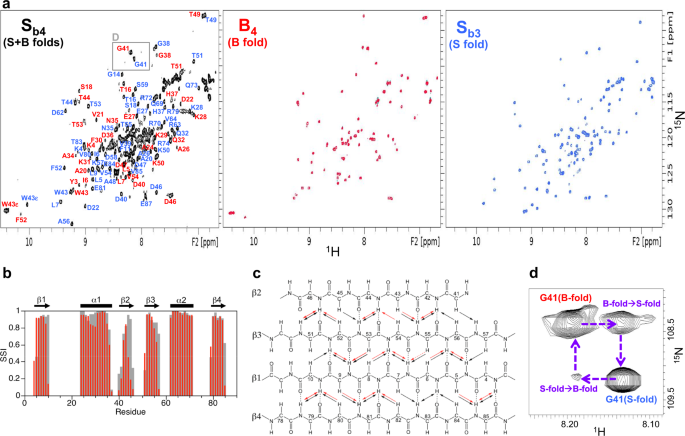
We used the NMR structure of Sb3to design a point mutation, tyrosine 5 to leucine (Y5L), that would stabilize the embedded B-fold without compromising native contacts in the S-fold (Supplementary Figure 10). This mutant was therefore expected to shift the population to the B-fold. Two mutants were prepared, a Y5L mutant of Sb3(denoted Sb4) and a Y5L mutant of B3(denoted B4).B4, is indeed more stable than B3(Fig.5a, Supplementary Table1).Assignment and structure determination of B4showed its topology to be identical to the parent B-fold (Fig.6b, c).At concentrations above 100 μM, B4displayed a tendency for weak self-association similar to that seen for B3。For Sb4, the HSQC spectrum exhibited approximately twice the number of amide cross-peaks relative to Sb3(Fig.7a), suggesting that S- and B-states were populated simultaneously. This was confirmed by the NMR assignment and also a comparison of the HSQC spectra for Sb4, B4, and Sb3。A significant fraction of the Sb4backbone amide signals (~50 peaks) closely matched those of B4, indicating the presence of a B-state (Supplementary Fig.12A–C).The close matching of these peaks is presumably because residues 1–56 in the B-state of Sb4are identical in sequence to B4。The largest amide shift perturbations between the B-state of Sb4and B4occur for residues proximal to the C-terminus of the B-fold, such as G41, where Sb4has additional residues and B4does not. Many of the Sb4signals also matched well with Sb3, although the degree of similarity was not as extensive as with B4(Supplementary Fig.12D–F).More significant amide chemical shift differences between the S-state of Sb4and Sb3are likely due to the Y5L mutation, which is a relatively large change located adjacent to the core. To resolve these ambiguities, backbone resonance assignments were made for the S-state of Sb4(Fig.7a, [https://doi.org/10.13018/BMR51719有关详细信息,请参阅“方法”部分)。Comparison of Sb4S-state assignments with Sb3indicated that most of the larger amide shift perturbations were in the β1 and β4 strands. Secondary shift analysis showed that the pattern of secondary structure elements for the S-state of Sb4is similar to that of Sb3(Fig.7b).Inter-proton NOE analysis indicated that the arrangement of the β-strands is also similar (Fig.7c).Together, these results show that Sb4populates both S- and B-folds approximately equally at 25 °C. Moreover, a ZZ-exchange spectrum demonstrated that the S- and B-states of Sb4are in slow conformational exchange on the NMR timescale (Fig.7d).
aTwo-dimensional1H–15N HSQC spectra of Sb4(left), B4(center), and Sb3(right) at 25 °C. In the Sb4spectrum, backbone amide resonance assignments are shown for the S-state (blue) and the B-state (red).bConfidence levels of ordered secondary structure regions for the S-state of Sb4at 30 °C (red) and for Sb3at 25 °C (gray) from chemical shifts using TALOS-N. The secondary structure elements of Sb3, determined from the three-dimensional structure, are shown above the plot.cSummary of long-range backbone NOEs from 3D15N-NOESY data for Sb3at 25 °C (black) and the S-state of Sb4at 30 °C (red).dGly41 region of the 300 ms ZZ-exchange spectrum for Sb4at 25 °C, showing exchange peaks between the S- and B-states.
Finally, we designed a mutation of leucine 67 to arginine (L67R) in Sb4to destabilize the S-fold without changing the sequence of the embedded B-fold. The mutant is denoted as Sb5(Supplementary Fig.10).This was expected to shift the population to the B-fold. The 2D1H-15N HSQC spectrum of Sb5indicates that the L67R mutation does indeed destabilize the S-fold, with the loss of S-type amide cross-peaks and the concurrent appearance of a new set of signals indicating a switch to a B-fold. The superposition of the spectrum of Sb5with that of B4shows that the new signals in Sb5largely correspond with the spectrum of B4(Supplementary Fig.13).Thus, the L67R mutation shifts the equilibrium from the S-fold to the B-fold. The additional signals (~25–30) in the central region of the HSQC spectrum that are not detected in B4are presumably due to the disordered C-terminal tail of Sb5。The C-terminal tail of Sb5does not appear to interact extensively with the B-fold, as evidenced by few changes in chemical shifts or peak intensities in the B-region of Sb5compared with B4。
Structural comparison of Sb3and B4
The aligned amino acids 1–56 of Sb3and B4have 98% sequence identity, the only difference being an L5Y mutation in Sb3(Fig.6a).The global folds of Sb3and B4have large-scale differences, however (Fig.8a, Supplementary Fig.4).The β1-strands, while similar in length, are in opposite directions in Sb3and B4。The β1-strand forms a parallel-stranded interaction with β4 in B4, but an antiparallel interaction with the corresponding β3-strand in Sb3。Whereas residues 9-20 form the 6-residue β1–β2 turn and the 6-residue β2-strand of B4, these same amino acids constitute the end of β1 and 10 residues of the largely disordered β1–α1 loop in Sb3。The remainder of the B-region is topologically similar, with the α1/β3/β4 structure in B4matching the α1/β2/β3 structure in Sb3。Overall, however, the order of H-bonding in the 4-stranded β-sheets is quite different, with β2β3β1β4 in Sb3and β3β4β1β2 in B4。
aMain chain comparisons. (Left panel) CS-Rosetta structure of B4with secondary structure elements color coded. (Right panel) Corresponding color-coded regions mapped onto the CS-Rosetta structure of Sb3, showing changes in backbone conformation. Regions outside the 56 amino acid sequence of B4are shown in wheat.bSide chain comparisons. (Left panel) Residues contributing to the core of B4from α1/β3/β4 (yellow), and from other regions (cyan). The non-α1/β2/β3 core residues from Sb3(pink) do not overlap with the B4core (see text for further details). (Right panel) Residues contributing to the core of Sb3from α1/β2/β3 (yellow), and the other participating core residues (pink). The non-α1/β2/β3 core residues from B4are also shown (cyan).. The single L5Y amino acid difference between B4and Sb3is highlighted.
The main core residues of B4consist of Y3, L5, L7, and L9 from β1, A26, F30, and A34 from α1, and F52 and V54 from β4 (Fig.8b).In Sb3, the topologically equivalent regions of the core are A26, F30, and A34 from α1, and F52 and V54 from β3. Residues Y5, L7, and L9 from the β1 strand of Sb3also form part of the core, but with different packing from B4due to the reverse orientation of β1. Residues A12 and A20, which contribute to the periphery of the core in B4, are solvent accessible in the β1-α1 loop of Sb3。Most of the remaining core residues of Sb3come from outside of the B-region and include amino acids from β3 (A56), α2 (V64, L67, A68, L71), and β4 (V80 and I82).
Energetics of unfolding for B3/Sb3, B4/Sb4, and Sb5
Far-UV CD spectra were measured for B3, B4, Sb3, Sb4, and Sb5and their thermal unfolding profiles were determined by measuring ellipticity at 222 nm versus temperature (Fig.5, Supplementary Fig.10, Supplementary Table1).As described above, the predominant form of Sb3is an S-fold. CD and NMR analyses show that B3is predominantly a B-fold with a ∆Gfoldingof −1.2 kcal/mol at 25 °C58。From the NMR analysis, it appears that the B-fold is in equilibrium with putatively dimeric states. This creates a situation in which the B-fold is both temperature-dependent and concentration-dependent. The predominant form at 5 °C and ≤18 µM is the B-fold, however. The low stability and concentration-dependent behavior of B3may indicate that some propensity for S-type conformations could persist in the 56-residue protein.
Sb4has a temperature unfolding profile very similar to Sb3(Fig.5) even though both S- and B- are approximately equally populated at 25 °C in Sb4(Fig.7).This shows that the Y5L mutation results in two folds that are almost isoenergetic and both thermodynamically stable relative to the unfolded state. Further, because S- and B-folds are in equilibrium and approximately equally populated, the free energy of switching to the B-fold from the S-fold (∆GB-fold/S-fold) is ~0 kcal/mol at 25 °C. The switch equilibrium reflects the influence of the antagonistic B-fold on the S-fold population in Sb4, where the leucine at residue 5 helps stabilize the alternative B-state at the expense of the S-state. Thermal denaturation by CD shows that B4has a ∆Gfolding = −4.1 kcal/mol at 25 °C58。The thermal unfolding profile of Sb5shows a low-temperature transition with a midpoint ~10 °C and a major transition with a midpoint of ~60 °C (Fig.5b).The NMR analysis indicates that the major transition is unfolding of the B-fold. Thus, the arginine at 67 in Sb5makes the B-fold more favorable by making the S-fold unfavorable, consistent with the change in population from mixed to B-fold observed by NMR.
Protease inhibition
The Sb3protein is closely related to S’I but lacks inhibitor function because C-terminal amino acids were changed in the design of the switch. It can be converted into a protease inhibitor, however, by altering C-terminal amino acids VTE to DKLYRAL. This mutant is denoted Sb3I. Sb3and Sb3I appear similar in structure by CD analysis (Supplementary Fig.10).The KIfor Sb3I with the engineered subtilisin was determined to be 50 nM (Supplementary Table1).
IgG binding
Binding to IgG was determined for B3and Sb3I (Supplementary Table1).B3and Sb3I bound to IgG Sepharose withKD ≤ 1 µM and 10 µM, respectively. Presumably, Sb3I has significant IgG-binding activity because the α1β3 IgG binding surface of the B-fold is largely preserved in the S-fold. Thus, Sb3I is a dual-function protein with both IgG-binding and protease inhibitor functions (Fig.2f).
Discussion
The entire network of intersecting pathways between the S-, A-, and B-folds is summarized in Fig.9。The first node on the pathway is a functional switch from RNA binding protein to protease inhibitor without a fold switch. The α/β plait is a common fold, and proteins with this basic topology include many different functions42。工程SI和S“节点说明protease inhibitor function can arise in the α/β plait topology with a few mutations. Replacing only C-terminal amino acids in the S6 protein creates interaction with the substrate binding cleft of the protease (Fig.2a, b).This C-terminal interaction plus adventitious contact between the β-sheet surface of the α/β plait and two α-helices in the protease result in protease inhibition in the 50 nM range. Based on the structure of S6 in the 30S complex, the C-terminal modification may not have major effects on binding interactions with ribosomal RNA and the S15 protein (Fig.2a)43。Thus, the transition from RNA binding protein to protease inhibitor likely is uninterrupted. An insertion in the β1–α1 loop and a deletion β2–β3 loop in the SI-inhibitor creates a topology that more closely resembles natural prodomain-type inhibitors44,46,61and creates an α1β2β3 motif in the S’-fold that is similar to the α1β3β4 motif of the B-fold. This topological similarity brings the S’I closer to an intersection with the B-fold. Thus, SI and S’I nodes are both functional switches and branch points for switching the S-fold into the A- and B-folds, respectively.
Switches between stable folds can be induced by a single amino acid mutation or deleting/appending terminal sequence that stabilizes the S-fold. Blue denotes an S-fold, green a B-fold, and red an A-fold. Gray arrows connect proteins that have been reengineered without a fold switch. Sb4is observed with two folds simultaneously. The GA98 and GB98 structures are from PDB codes2LHCand2LHD(ref.32), respectively.
Engineering nodes at fold intersections required designing sequences that are compatible with native interactions in two different folds. We used simple rules to do this. The first rule was to align topologies rather than maximizing sequence similarities. Identifying a common topology can help determine a register that has fewer irreconcilable clashes. For example, topological alignment of the α1 helix of the SI fold and the α1 helix of the A-fold facilitated engineering the fold switch, because the regions flanking α1 of the SI-fold can encode two different fold motifs. When topological alignment is poor, as was the case with S- and B-folds, it was helpful to look for natural variations in the turns of the longer fold to create better alignment. Variation in loops and turns in a larger fold creates more freedom of design and a higher probability of switches. Once an alignment is chosen, the basic rule in resolving catastrophic clashes is to conserve original amino acids when possible. This reduces the uncertainties involved in computational design. The Rosetta energy function was not used to predict a favorable alignment but was important in evaluating mutations to resolve clashes once an alignment was chosen.
Selecting mutations compatible with two sets of native interactions required tradeoffs in the native state energetics of each individual fold5,11。A node may be produced in cases in which both alternative folds are stable relative to the unfolded state. Stability relative to the unfolded state (i.e. a state with little secondary structure) was determined by CD melting (Fig.5).It was informative to examine the stability of both short (56 residues) and longer forms of a putative node sequence. The independent stability of the G-fold can be determined in the short form without the antagonism from the S-fold that is present in the longer sequence. The stabilities of the A1and A2proteins are about −4 kcal/mol at 25 °C58compared to −5.6 kcal/mol for the native GAprotein56。The stabilities of B3and B4are −1.2 and −4.1 kcal/mol, respectively, at 25 °C58compared to −6.7 kcal/mol for the native GBprotein62。For the longer sequences, the ∆Gfoldingof Sa1and Sb3are −5.3 and −3.5 kcal/mol, respectively, at 25 °C58compared to −8.5 kcal/mol for the native S6 protein40。
然而在S-folds的情况下,energetic effects of the stable, embedded G-fold must also be considered. Since the equilibria between both folded states and the unfolded state are thermodynamically linked, the free energy of a switch to a G-fold from an S-fold (∆GG-fold/S-fold) is approximated by the difference in ∆Gfolding(∆∆Gfolding) between the short and long forms of a node protein. For example, based on ∆Gfoldingfor A1and Sa1, the predicted ∆GA-fold/S-foldof Sa1is 1.3 kcal/mol. This is consistent with the structure of the predominant S-fold determined by NMR but also with the small population of 3α fold suggested by weak HSA-binding. From the thermal denaturation profiles of B3and Sb3, the predicted ∆GB-fold/S-foldof Sb3is 2.3 kcal/mol, a value consistent with the stable S-fold observed in NMR experiments. The Sb3sequence is also approaching a critical point, however. A substitution in Sb3that stabilizes the B-fold (Y5L) shifts the equilibrium of Sb4to an approximately equal mixture of B- and S-folds. That is, ∆GB-fold/S-foldof Sb4is ~0 kcal/mol at 25 °C. One further substitution that destabilizes the S-fold (L67R) shifts the population of Sb5to a stable B-fold (∆GB-fold/S-fold ≤ −5 kcal/mol) (Fig.9).
The existence of nodes between folds has implications for the evolution of new functions. In the case of the S/A node, all contact amino acids for HSA exist within the S-fold of the protease inhibitor Sa2I albeit in a cryptic topology. Deletion of amino acids 67–99 (A2) results in loss of inhibitor function and a fold switch from α/β plait to 3α. Acquisition of HSA binding activity (KD < 1 µM) results from unmasking the cryptic HSA binding amino acids via the fold switch (Fig.2e).This level of binding affinity could be biologically relevant since the concentration of HSA in serum is >500 µM63。In the case of the S/B node, the α1β3 motif contains all IgG contact amino acids and Sb3I has some affinity for both IgG (KD = 10 µM) and protease (KI = 50 nM). In this case, the Y5L mutation (Sb4) or a deletion of 57–91 (B4) causes a fold switch from α/β plait to the β-grasp and results in tighter IgG binding (KD ≤ 1 µM) (Fig.2f).This level of binding affinity could also be biologically relevant since the concentration of IgG in serum is >50 µM (or >100 µM Fc binding sites)64。We have previously shown that an A-fold with HSA binding function can be switched to a B-fold with IgG-binding function via single amino acid substitutions that switch the folds and unmask cryptic contact amino acids for the two ligands29,32。
总之,可以连接三个公司mmon folds in a network of high-identity nodes that form critical points between two folds. As in other complex systems, a small change in a protein near a critical point can have a “butterfly effect” on how the folds are populated. This property of the protein folding code means that proteins with multiple folds and functions can exist in highly identical amino acid sequences. This suggests that the evolution of new folds and functions sometimes can follow uninterrupted mutational pathways.
Methods
Mutagenesis, protein expression and purification
诱变是使用Q5®Site-Directed Mutagenesis Kits (NEB). GAand GBvariants were cloned into a vector (pH0720) encoding the sequence:
MEAVDANSLA QAKEAAIKEL KQYGIGDKYI KLINNAKTVE GVESLKNEIL KALPTEGSGN TIRVIVSVDK AKFNPHEVLG IGGHIVYQFK LIPAVVVDVP ANAVGKLKKM PGVEKVEFDH QYRGL
as an N-terminal fusion domain56。Cell growth was carried out by auto-induction29,65。Cells were harvested by centrifugation at 3750 × gfor 20 min and lysed by sonication on ice in 0.1 M KPi, pH 7.2. Cellular debris was pelleted by centrifugation at 10,000 × gfor 15 min. Supernatant was clarified by centrifugation at 45,000 × gfor 30 min. Proteins were purified using a second generation of the affinity-cleavage tag system employed previously to purify switch proteins29,66。The second-generation tag results in high-level soluble expression of the switch proteins and also enables the capture of the fusion protein by binding tightly to an immobilized processing protease via the C-terminal EFDHQYRGL sequence. Loading and washing were at 5 mL/min for a 5 mLIm-Prot列使用运行缓冲的20毫米KPi, pH值6.8。The amount of washing required for high purity depends on the stickiness of the target protein and how much of it is bound to the column. We typically wash with 10 column volumes (CV) of wash solution followed by 3 CV 0.5 M NaCl and then ~10 CV running buffer. This can be repeated as necessary. The 0.5 M NaCl shots are repeated until the amount of absorbance released with each high-salt shot becomes small and constant. All the high-salt solution is washed out before initiating the cleavage. The target protein was cleaved from theIm-Prot列通过注射15毫升的咪唑溶液(0。1 mM) at 1 mL/min, 22 °C. The cleaved protein typically elutes as a sharp peak in 2–3 CV. The purified protein was then concentrated to 0.2–0.3 mM, as required for NMR analysis. The columns were regenerated by injecting 15 mL of 0.1 N H3PO4(0.227 mL concentrated phosphoric acid (85%) per 100 mL) at a flow rate of ~1 CV/min. The wash solution was neutralized immediately after stripping. The purification system is available from Potomac Affinity Proteins.
Protease inhibitor proteins were purified by binding toIm-Protmedia and then stripping off the purified inhibitor with 0.1 N H3PO4。Samples were then immediately neutralized by adding 1/10 volume 1 M K2HPO4。
Rosetta calculations
Rosetta energies of all designed structures were generated using the Slow Relax routine54。1000 decoys were calculated for each design. PDB coordinates and energy parameters for the lowest energy decoy for each design are included as supplemental files.
Circular dichroism (CD)
CD measurements were performed in 100 mM KPi, pH 7.2 with a Jasco spectropolarimeter, model J-1100 with a Peltier temperature controller. Quartz cells with path lengths of 0.1 and 1 cm were used for protein concentrations of 3 and 30 µM, respectively. The ellipticity results were expressed as mean residue ellipticity, [θ], deg cm2 dmol−1。Ellipticities at 222 nm were continuously monitored at a scanning rate of 0.5°/min. Reversibility of denaturation was confirmed by comparing the CD spectra at 20 °C before melting and after heating to 100 °C and cooling to 20 °C.
Measuring HSA and IgG binding affinity
Affinity of proteins to HSA and IgG was determined by their retention on the immobilized ligands. HSA and rabbit IgG were immobilized by reaction with NHS-activated Sepharose 4 Fast Flow (Cytiva) according to the manufacturer’s instructions. The concentration of immobilized HSA was 100 µM. The concentration of immobilized IgG was 50 µM (i.e. 100 µM Fc binding sites). Generally, 0.2 mL of a 5 µM solution of the test protein was injected into a 5 mL column at a flow rate of 0.5 mL/min. Determination of binding affinity assumes that binding is in rapid equilibrium such that the elution volume is proportional to the fraction of test protein bound to 100 µM of binding sites. Proteins that are completely retained after 20 column volumes (CV) are assessed to haveKD ≤ 1 µM. Completely retained proteins are stripped from the column with 0.1 N H3PO4at the end of the run.
Measuring protease inhibition
Competitive inhibition constants (KI) were determined using the fluorogenic peptide substrate QEEYSAM-AMC (7-amino-4-methylcoumarin) purchased from AnaSpec Inc. and a highly specific, engineered protease known as RASProtease(I)49。Competitive inhibition constants (KI) were measured by determining theKM(apparent)in the presence of 0, 50, and 100 nM of each inhibitor protein. The reactions were carried out in 100 mM KPi, 10 mM imidazole, 0.005% tween-20, pH 7.0 at 25 °C with 1 nM RASProtease(I). The QEEYSAM-AMC concentrations used to determineKMandKM(apparent)were 0.1, 0.5, 1, 2, 5, and 10 µM. Initial rates were determined with a BioTek Synergy MT fluorescence microplate reader (Ex: 360/40, Em: 460/40) by measuring the release of the fluorescent AMC group via hydrolysis of the amide bond. Highly pure (≥98%) protease and inhibitor proteins were used for all kinetic experiments.
NMR spectroscopy
Isotope-labeled samples were prepared at 0.2–0.3 mM concentrations in 100 mM potassium phosphate buffer (pH 7.0) containing 5% D2O. NMR spectra were collected using Topspin3.6.1 software on Bruker AVANCE III 600 and 900 MHz spectrometers fitted with Z-gradient1H/13C/15N三重共振冷冻器。标准双和triple resonance experiments (HNCACB, CBCA(CO)NH, HNCO, HN(CA)CO, and HNHA) were utilized to determine main chain NMR assignments. Inter-proton distances were obtained from 3D15N-edited NOESY and 3D13C-edited NOESY spectra with a mixing time of 150 ms. NmrPipe67was used for data processing and analysis was done with Sparky68。Two-dimensional {1H}-15N steady-state heteronuclear NOE experiments were acquired with a 5 s relaxation delay between experiments. Errors in heteronuclear NOEs were estimated based on the background noise level. Chemical shift perturbations were calculated using Δδtotal = ((WHΔδH)2 + (WNΔδN)2)1/2, whereWHis 1,WNis 0.2, and ΔδHand ΔδNrepresent1H and15N chemical shift changes, respectively. For PRE experiments on Sb1, single-site cysteine mutant samples were incubated with 10 equivalents of (1-oxyl-2,2,5,5-tetramethylpyrroline-3-methyl) methanethiosulfonate (MTSL), Santa Cruz Biotechnology) at 25 °C for 1 h and completion of labeling was confirmed by MALDI mass spectrometry. Control samples were reduced with 10 equivalents of sodium ascorbate. Backbone amide peak intensities of the oxidized and reduced states were analyzed using Sparky. Three-dimensional structures were calculated with CS-Rosetta3.2 using experimental backbone15N,1HN,1Hα13Cα,13Cβ, and13CO chemical shift restraints and were either validated by comparison with experimental backbone NOE patterns (A1, B1, B4, Sb1) or directly employed interproton NOEs (Sa1, Sb2) or PREs (Sb1) as additional restraints. One thousand CS-Rosetta structures were calculated from which the 10 lowest energy structures were chosen. For Sb3, CS-Rosetta failed to converge to a unique low-energy topology, producing an approximately even mixture of S- and B-type folds despite the chemical shifts and NOE pattern indicating an S-fold. In this case, CNS1.169was employed to determine the structure56, including backbone dihedral restraints from chemical shift data using TALOS-N70。The backbone resonances for the S-state of Sb4were assigned using triple resonance methods as above, under conditions where the S-state is more favorably populated (30 °C, 100 mM KPi, 200 mM sodium chloride, pH 7.0). Amide assignments were then transferred to the two-dimensional1H-15N HSQC spectrum of Sb4at 25 °C in 100 mM KPi, pH 7.0. Inter-proton NOEs for the S-state of Sb4were obtained at the 30 °C/high salt condition, employing a 3D15N-edited NOESY spectrum with a 150 ms mixing time. A two-dimensional ZZ-exchange1H–15N HSQC spectrum was recorded on Sb4using a mixing time of 300 ms (25 °C, 100 mM KPi, pH 7.0)71,72。Protein structures were displayed and analyzed utilizing PROCHECK-NMR73, MOLMOL74and PyMol (Schrodinger)55。
Reporting summary
Further information on research design is available in theNature Portfolio Reporting Summarylinked to this article.
Data availability
The NMR structures generated in this study have been deposited in the PDB: [https://doi.org/10.2210/pdb7MN1/pdb]; [https://doi.org/10.2210/pdb7MQ4/pdb]; [https://doi.org/10.2210/pdb7MN2/pdb]; [https://doi.org/10.2210/pdb7MP7/pdb]; [https://pdb-dev.wwpdb.org/entry.html?PDBDEV_00000083]; [https://pdb-dev.wwpdb.org/entry.html?PDBDEV_00000084]; [https://pdb-dev.wwpdb.org/entry.html?PDBDEV_00000085]. NMR Assignments have been deposited in the BMRB: [https://doi.org/10.13018/BMR30901]; [https://doi.org/10.13018/BMR30902]; [https://doi.org/10.13018/BMR30904]; [https://doi.org/10.13018/BMR30905]; [https://doi.org/10.13018/BMR50907]; [https://doi.org/10.13018/BMR50909]; [https://doi.org/10.13018/BMR50910]; [https://doi.org/10.13018/BMR51719]. The structures referenced in this paper are publicly available in the PDB: [https://doi.org/10.2210/pdb1FKA/pdb]; [https://doi.org/10.2210/pdb2VDB/pdb]; [https://doi.org/10.2210/pdb1FCC/pdb]; [https://doi.org/10.2210/pdb6UAO/pdb]; [https://doi.org/10.2210/pdb2LHC/pdb]; [https://doi.org/10.2210/pdb1RIS/pdb]. Source data are provided with this paper. Design models are provided as files in the source data.Source dataare provided with this paper.
References
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold.Nature596583 - 589 (2021).
Baek, M. et al. Accurate prediction of protein structures and interactions using a three-track neural network.Science373, 871–876 (2021).
Huang, P. S., Boyken, S. E. & Baker, D. The coming of age of de novo protein design.Nature537, 320–327 (2016).
Ambroggio, X. I. & Kuhlman, B. Design of protein conformational switches.Curr. Opin. Struct. Biol.16, 525–530 (2006).
Bryan, P. N. & Orban, J. Proteins that switch folds.Curr. Opin. Struct. Biol.20, 482–488 (2010).
Dishman, A. F. et al. Evolution of fold switching in a metamorphic protein.Science371, 86–90 (2021).
Wei, K. Y. et al. Computational design of closely related proteins that adopt two well-defined but structurally divergent folds.Proc. Natl Acad. Sci. USA117, 7208–7215 (2020).
Anderson, W. J., Van Dorn, L. O., Ingram, W. M. & Cordes, M. H. Evolutionary bridges to new protein folds: design of C-terminal Cro protein chameleon sequences.Protein Eng. Des. Sel.24, 765–771 (2011).
Burmann, B. M. et al. An α helix to β barrel domain switch transforms the transcription factor RfaH into a translation factor.Cell150, 291–303 (2012).
Kulkarni, P. et al. Structural metamorphism and polymorphism in proteins on the brink of thermodynamic stability.Protein Sci.27, 1557–1567 (2018).
Dishman, A. F. & Volkman, B. F. Design and discovery of metamorphic proteins.Curr. Opin. Struct. Biol.74, 102380 (2022).
Alberstein, R. G., Guo, A. B. & Kortemme, T. Design principles of protein switches.Curr. Opin. Struct. Biol.72, 71–78 (2022).
Rackovsky, S. Nonlinearities in protein space limit the utility of informatics in protein biophysics.Proteins83, 1923–1928 (2015).
Chen, S. H., Meller, J. & Elber, R. Comprehensive analysis of sequences of a protein switch.Protein Sci.25, 135–146 (2016).
Li, W., Kinch, L. N., Karplus, P. A. & Grishin, N. V. ChSeq: A database of chameleon sequences.Protein Sci.24, 1075–1086 (2015).
Wolynes, P. G. Evolution, energy landscapes and the paradoxes of protein folding.Biochimie119, 218–230 (2015).
Holzgräfe, C. & Wallin, S. Smooth functional transition along a mutational pathway with an abrupt protein fold switch.Biophys. J.107, 1217–1225 (2014).
Scheraga, H. A. & Rackovsky, S. Homolog detection using global sequence properties suggests an alternate view of structural encoding in protein sequences.Proc. Natl Acad. Sci. USA111, 5225–5229 (2014).
Ha, J. H. & Loh, S. N. Protein conformational switches: from nature to design.Chemistry18, 7984–7999 (2012).
Yadid, I., Kirshenbaum, N., Sharon, M., Dym, O. & Tawfik, D. S. Metamorphic proteins mediate evolutionary transitions of structure.Proc. Natl Acad. Sci. USA107, 7287–7292 (2010).
Lichtarge, O. & Wilkins, A. Evolution: a guide to perturb protein function and networks.Curr. Opin. Struct. Biol.20, 351–359 (2010).
Rollins, N. J. et al. Inferring protein 3D structure from deep mutation scans.Nat. Genet.51, 1170–1176 (2019).
Sikosek, T., Chan, H. S. & Bornberg-Bauer, E. Escape from Adaptive Conflict follows from weak functional trade-offs and mutational robustness.Proc. Natl Acad. Sci. USA109, 14888–14893 (2012).
Chen, N., Das, M., LiWang, A. & Wang, L. P. Sequence-based prediction of metamorphic behavior in proteins.Biophys. J.119, 1380–1390 (2020).
Porter, L. L. & Looger, L. L. Extant fold-switching proteins are widespread.Proc. Natl Acad. Sci. USA115, 5968–5973 (2018).
Bedford, J. T., Poutsma, J., Diawara, N. & Greene, L. H. The nature of persistent interactions in two model β-grasp proteins reveals the advantage of symmetry in stability.J. Comput. Chem.42, 600–607 (2021).
Sykes, J., Holland, B. R. & Charleston, M. A. A review of visualisations of protein fold networks and their relationship with sequence and function.Biol. Rev. Camb. Philos. Soc.https://doi.org/10.1111/brv.12905(2022).
Ambroggio, X. I. & Kuhlman, B. Computational design of a single amino acid sequence that can switch between two distinct protein folds.J. Am. Chem. Soc.128, 1154–1161 (2006).
Alexander, P. A., He, Y., Chen, Y., Orban, J. & Bryan, P. N. A minimal sequence code for switching protein structure and function.Proc. Natl Acad. Sci. USA106, 21149–21154 (2009).
Davey, J. A., Damry, A. M., Goto, N. K. & Chica, R. A. Rational design of proteins that exchange on functional timescales.Nat. Chem. Biol.13, 1280–1285 (2017).
He, Y., Chen, Y., Alexander, P., Bryan, P. N. & Orban, J. NMR structures of two designed proteins with high sequence identity but different fold and function.Proc. Natl Acad. Sci. USA105, 14412–14417 (2008).
He, Y., Chen, Y., Alexander, P. A., Bryan, P. N. & Orban, J. Mutational tipping points for switching protein folds and functions.Structure20, 283–291 (2012).
Falkenberg, C., Bjorck, L. & Akerstrom, B. Localization of the binding site for streptococcal protein G on human serum albumin. Identification of a 5.5-kilodalton protein G binding albumin fragment.Biochemistry31, 1451–1457 (1992).
Frick, I. M. et al. Convergent evolution among immunoglobulin G-binding bacterial proteins.Proc. Natl Acad. Sci. USA89, 8532–8536 (1992).
Myhre, E. B. & Kronvall, G. Heterogeneity of nonimmune immunoglobulin Fc reactivity among gram-positive cocci: description of three major types of receptors for human immunoglobulin G.Infect. Immun.17, 475–482 (1977).
Reis, K. J., Ayoub, E. M. & Boyle, M. D. P., Streptococcal Fc receptors. II. Comparison of the reactivity of a receptor from a group C streptococcus with staphylococcal protein A.J. Immunol.132, 3098–3102 (1984).
Lindberg, M. O., Haglund, E., Hubner, I. A., Shakhnovich, E. I. & Oliveberg, M. Identification of the minimal protein-folding nucleus through loop-entropy perturbations.Proc. Natl Acad. Sci. USA103, 4083–4088 (2006).
Haglund, E., Lindberg, M. O. & Oliveberg, M. Changes of protein folding pathways by circular permutation. Overlapping nuclei promote global cooperativity.J. Biol. Chem.283, 27904 - 27915(2008)。
Haglund, E. et al. The HD-exchange motions of ribosomal protein S6 are insensitive to reversal of the protein-folding pathway.Proc. Natl Acad. Sci. USA106, 21619–21624 (2009).
Haglund, E. et al. Trimming down a protein structure to its bare foldons: spatial organization of the cooperative unit.J. Biol. Chem.287, 2731–2738 (2012).
Lindahl, M. et al. Crystal structure of the ribosomal protein S6 fromThermus thermophilus。EMBO J.13, 1249–1254 (1994).
Day, R., Beck, D. A., Armen, R. S. & Daggett, V. A consensus view of fold space: combining SCOP, CATH, and the Dali Domain Dictionary.Protein Sci.12, 2150–2160 (2003).
Schluenzen, F. et al. Structure of functionally activated small ribosomal subunit at 3.3 angstroms resolution.Cell102, 615–623 (2000).
Gallagher, T. D., Gilliland, G., Wang, L. & Bryan, P. The prosegment-subtilisin BPN’ complex: crystal structure of a specific foldase.Structure3, 907–914 (1995).
Tangrea, M. A. et al. Stability and global fold of the mouse prohormone convertase 1 pro-domain.Biochemistry40, 5488–5495 (2001).
Tangrea, M. A., Bryan, P. N., Sari, N. & Orban, J. Solution structure of the pro-hormone convertase 1 pro-domain fromMus musculus。J. Mol. Biol.320, 801–812 (2002).
Sari, N. et al. Hydrogen-deuterium exchange in free and prodomain-complexed subtilisin.Biochemistry46, 652–658 (2007).
Orengo, C. A. & Thornton, J. M. Alpha plus beta folds revisited: some favoured motifs.Structure1, 105–120 (1993).
Chen, Y. et al. Engineering subtilisin proteases that specifically degrade active RAS.Commun. Biol.4, 299 (2021).
Lejon, S., Frick, I. M., Bjorck, L., Wikstrom, M. & Svensson, S. Crystal structure and biological implications of a bacterial albumin binding module in complex with human serum albumin.J. Biol. Chem.279, 42924–42928 (2004).
Sauer-Eriksson, A. E., Keywegt, G. J., Uhlen, M. & Jones, T. A. Crystal structure of the C2 fragment of streptococcal protein G in complex with the Fc domain of human IgG.Structure3, 265–278 (1995).
Alexander, P. A., Rozak, D. A., Orban, J. & Bryan, P. N. Directed evolution of highly homologous proteins with different folds by phage display: implications for the protein folding code.Biochemistry44, 14045–14054 (2005).
Alexander, P. A., He, Y., Chen, Y., Orban, J. & Bryan, P. N. The design and characterization of two proteins with 88% sequence identity but different structure and function.Proc. Natl Acad. Sci. USA104, 11963–11968 (2007).
Leaver-Fay, A. et al. ROSETTA3: an object-oriented software suite for the simulation and design of macromolecules.Methods Enzymol.487, 545–574 (2011).
Delano, W. L. The PyMOL Molecular Graphics System (DeLano Scientific, San Carlos, CA, 2002).
He, Y. et al. Structure, dynamics, and stability variation in bacterial albumin binding modules: implications for species specificity.Biochemistry45, 10102–10109 (2006).
Shen, Y. et al. De novo structure generation using chemical shifts for proteins with high-sequence identity but different folds.Protein Sci.19, 349–356 (2010).
Chen, Y. et al. Rules for designing protein fold switches and their implications for the folding code. Preprint atbioRxivhttps://doi.org/10.1101/2021.05.18.444643(2021).
Rozak, D. A., Orban, J. & Bryan, P. N. G148-GA3: a streptococcal virulence module with atypical thermodynamics of folding optimally binds human serum albumin at physiological temperatures.Biochim. Biophys. Acta1753, 226–233 (2005).
He, Y., Chen, Y., Rozak, D. A., Bryan, P. N. & Orban, J. An artificially evolved albumin binding module facilitates chemical shift epitope mapping of GA domain interactions with phylogenetically diverse albumins.Protein Sci.16, 1490–1494 (2007).
He, Y. et al. Solution NMR structure of a sheddase inhibitor prodomain from the malarial parasitePlasmodium falciparum。Proteins80, 2810–2817 (2012).
Alexander, P., Fahnestock, S., Lee, T., Orban, J. & Bryan, P. Thermodynamic analysis of the folding of the Streptococcal protein G IgG-binding domains B1 and B2: why small proteins tend to have high denaturation temperatures.Biochemistry31, 3597–3603 (1992).
Chien, S.-C., Chen, C.-Y., Lin, C.-F. & Yeh, H.-I. Critical appraisal of the role of serum albumin in cardiovascular disease.Biomark. Res.5, 31 (2017).
Gonzalez-Quintela, A. et al. Serum levels of immunoglobulins (IgG, IgA, IgM) in a general adult population and their relationship with alcohol consumption, smoking and common metabolic abnormalities.Clin. Exp. Immunol.151, 42–50 (2008).
Studier, F. W. Protein production by auto-induction in high density shaking cultures.Protein Expr. Purif.41, 207–234 (2005).
Ruan, B., Fisher, K. E., Alexander, P. A., Doroshko, V. & Bryan, P. N. Engineering subtilisin into a fluoride-triggered processing protease useful for one-step protein purification.Biochemistry43, 14539–14546 (2004).
Delaglio, F. et al. NMRPipe: a multidimensional spectral processing system based on UNIX pipes.J. Biomol. NMR6, 277–293 (1995).
Goddard, D. & Kneller, D. G. SPARKY 3 Vol. 3 (University of California, San Francisco, 2004).
Brunger, A. T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination.Acta Crystallogr. D (Biol. Crystallogr.)54, 905–921 (1998).
Shen, Y. & Bax, A. Protein backbone and sidechain torsion angles predicted from NMR chemical shifts using artificial neural networks.J. Biomol. NMR56, 227–241 (2013).
Farrow, N. A., Zhang, O., Forman-Kay, J. D. & Kay, L. E. A heteronuclear correlation experiment for simultaneous determination of15N longitudinal decay and chemical exchange rates of systems in slow equilibrium.J. Biomol. NMR4, 727–734 (1994).
Montelione, G. T. & Wagner, G. 2D Chemical exchange NMR spectroscopy by proton-detected heteronuclear correlation.J. Am. Chem. Soc.111, 3096–3098 (1989).
Laskowski, R. A., Rullmann, J. A., MacArthur, M. W., Kaptein, R. & Thornton, J. M. AQUA and PROCHECK-NMR: Programs for checking the quality of protein structures solved by NMR.J. Biomol. NMR8, 477–486 (1996).
Koradi, R., Billeter, M. & Wuthrich, K. MOLMOL: a program for display and analysis of macromolecular structures.J. Mol. Graph. Model.14, 51–55 (1996).
Acknowledgements
This work was supported by National Institutes of Health Grant GM62154 (to P.B. and J.O.) and 5R44GM126676 (to P.B.). The NMR facility is supported by the University of Maryland, the National Institute of Standards and Technology, and a grant from the W. M. Keck Foundation. We also thank Drs. Nese Sari and Louisa Wu for critically reading the manuscript and for many thoughtful comments. Mention of commercial products does not imply recommendation or endorsement by NIST.
Author information
Authors and Affiliations
Contributions
Protein design: Yw.C., B.R., E.C., J.O., P.B.; Performed thermodynamic and binding analyses: B.R., Yw.C., D.M., R.S., P.B.; Performed dynamic light scattering experiments: T.G.; Performed NMR experiments/structural analysis: Y.H., Yh.C., T.S., T.K., J.O.; Wrote the paper: J.O. (NMR and structural analysis), Yw.C., B.R., P.B. (remaining sections).
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communicationsthanks the anonymous, reviewer(s) for their contribution to the peer review of this work.Peer reviewer reportsare available.
Additional information
Publisher’s noteSpringer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open AccessThis article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visithttp://creativecommons.org/licenses/by/4.0/。
About this article
Cite this article
Ruan, B., He, Y., Chen, Y.et al.Design and characterization of a protein fold switching network.Nat Commun14, 431 (2023). https://doi.org/10.1038/s41467-023-36065-3
Received:
Accepted:
Published:
DOI:https://doi.org/10.1038/s41467-023-36065-3
Comments
By submitting a comment you agree to abide by ourTermsandCommunity Guidelines。If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.